A kind of biotechnology involving manipulation of DNA is:
1. DNA replication
2. genetic engineering
3. denaturation
4. renaturation
2. DNase I
3. RNase
4. Hind II
Match the following enzymes with their functions:
|
Column-I |
Column-II |
||
|
(a) |
Restriction endonuclease |
(i) |
joins the DNA fragments |
|
(b) |
Exonuclease |
(ii) |
extends primers on genomic DNA template |
|
(c) |
DNA ligase |
(iii) |
cuts DNA at specific position |
|
(d) |
Tag polymerase |
(iv) |
removes nucleotides from the ends of DNA |
Select the correct option from the following:
1. (a) - (iii), (b)- (i), (c)- (iv), (d)-(ii)
2. (a) - (iii), (b)- (iv), (c)- (i), (d)-(ii)
3. (a) - (iv), (b)- (iii), (c)- (i), (d)-(ii)
4. (a) - (ii), (b)- (iv), (c)- (i), (d)-(iii)
| 1. | It cuts DNA fragments into smaller pieces to ensure proper replication. |
| 2. | It joins DNA fragments together by forming phosphodiester bonds between them. |
| 3. | It ensures that the foreign DNA is not rejected by the host organism. |
| 4. | It prevents foreign DNA from replicating outside of the host’s genome. |
Complete the analogy:
Molecular scissor : Restriction endonuclease : : Molecular glue : X
| 1. | Alkaline phosphatase |
| 2. | Acid phosphatase |
| 3. | Taq polymerase |
| 4. | DNA ligase |
| 1. | They protect the DNA fragments from exonucleases. |
| 2. | They prevent the insertion of DNA fragments into the vector. |
| 3. | They ensure that the DNA fragments cannot be re-ligated by DNA ligase. |
| 4. | They allow fragments with identical sticky ends to hydrogen bond and facilitate ligation. |
If an enzyme catalyses the removal of nucleotides from the ends of DNA, then it should be called as:
1. endonuclease
2. exonuclease
3 ligase
4. reverse transcriptase
| 1. | The first letter represents the genus, the next two letters represent the species, the strain is indicated by an additional letter, and Roman numerals indicate the order of discovery. |
| 2. | The first letter represents the genus, the next two letters represent the species, and Roman numerals indicate the recognition sequence. |
| 3. | The first three letters represent the bacterial species, followed by the strain name and Roman numerals indicating the recognition sequence. |
| 4. | The entire name of the bacterial strain is used, followed by Roman numerals indicating the enzyme's function. |
What is true A,B and C in the given diagrammatic representation of rDNA technology ?
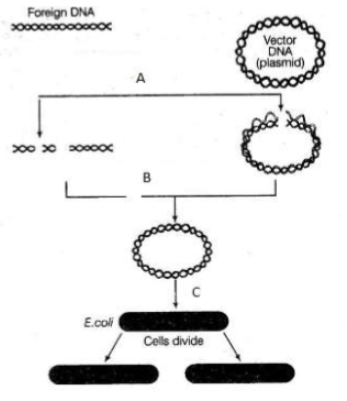
| I: | At A same restriction enzyme is used to cut both foreign and vector DNA |
| II: | The enzyme used at B is DNA ligase |
| III: | Step C can be called as transformation |
| 1. | I and II only |
| 2. | I and III only |
| 3. | II and III only |
| 4. | I,II and III |
| 1. | Because plasmids contain antibiotic resistance genes that trigger replication. |
| 2. | Because plasmids are naturally resistant to restriction enzymes, making replication possible. |
| 3. | Because plasmids possess an origin of replication that allows them to replicate autonomously. |
| 4. | Because plasmids are self-sustaining entities that can exist outside the nucleus. |






